
informatik



In order to query for information associated with the PHP domain, it is necessary to know the Pfam accession number of this domain. The graphical user interface provides a full text search to find this number.

|
The search frame provides the possibility to search the database with keywords for the accession number of entities. The results are presented in a table in the same frame and a double-click on the correct result inserts the entity into the query. The query can be completed using the selection buttons or manually. The complete query for species that have proteins with the PHP domain (Pfam acc: PF02811) and for biological processes of these proteins reads
TAX, GO WHERE PFAM:PF02811 RESTRICT TT:biological_process, SPECIES
This query combines queries for taxa and for GO terms. It limits the GO results to terms from the biological process ontology and the taxonomy results to species. The results are shown in tabs in a separate frame; a new tab is created for each table.
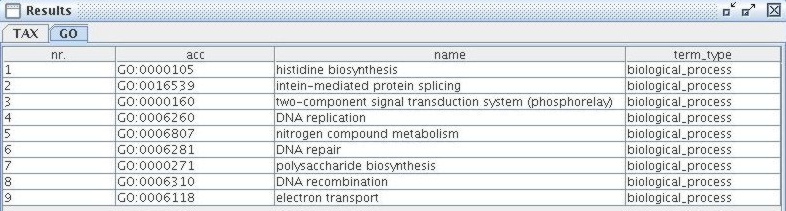
|
The tables contain all relevant data and provide the possibility to access the online source database with a pop-up menu. Both results are also highlighted in the respective tree views.
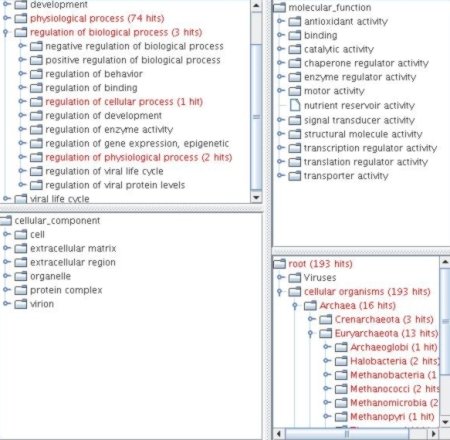
|
The four panes can be resized to view larger parts of one tree. The tree representation is very useful for browsing the hierarchy and locating specific nodes. A double-click on a node of a tree inserts this entity into the query field. This can be used to easily add a GO term to the query for further analysis. The query
GENE WHERE PFAM:PF02811 AND GO:GO:0006313
returns all proteins that have the PHP domain and are involved in "DNA recombination". The results are again added in a new tab to the results frame. It is possible to directly open a browser with the UniProt entry page for a protein from the list of results. Hovering over any tab with the mouse will display the query that returned this result as tooltip, i.e. the query is shown besides the mouse pointer. GOTaxExplorer allows to save single result sets to a tab-delimited text file. Furthermore, it is possible to delete single result tabs or to delete them all at once. GO results can contain detailed and more generic terms. GOTaxExplorer filters out generic terms if there are also descendants of these terms in the result set. However, it is possible to receive all terms. Another option allows to print all possible paths from result terms to the root.